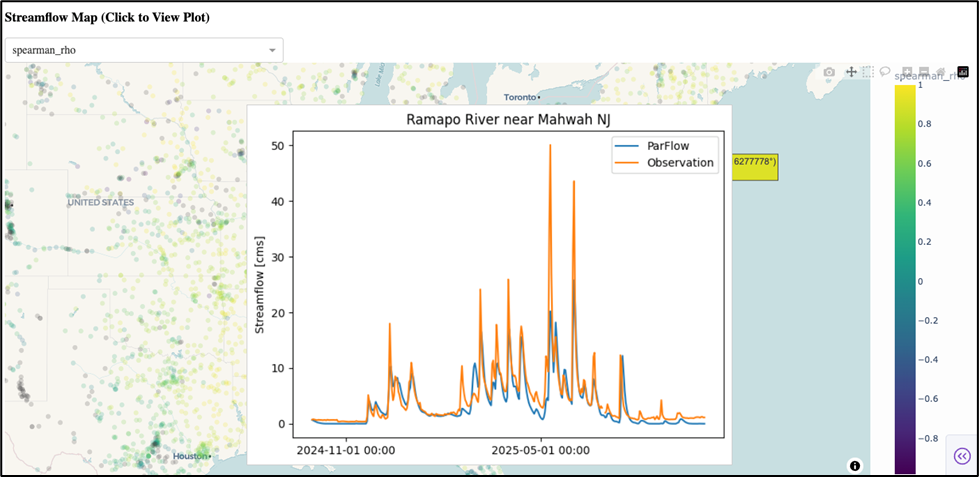
SPECFEM has been a cornerstone of computational seismology for over two decades. The Fortran-based software suite is used for seismic wave propagation simulations and adjoint tomography, accumulating thousands of citations per year across a broad user base. But the original codebase grew across three separate repositories (2D, 3D, and 3D Globe) with substantial redundancy and architecture-specific GPU kernels, meaning new features rarely made it to all variants.
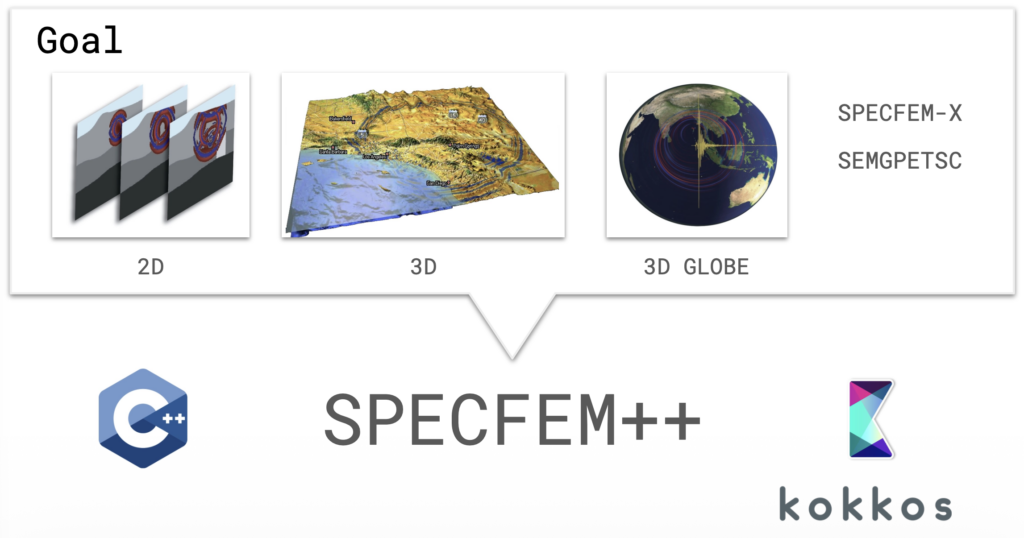
SPECFEM++ is the solution: a ground-up rewrite in C++ using Kokkos for performance portability across CPUs, NVIDIA GPUs (CUDA), and AMD GPUs (HIP) — all from a single codebase.

Expanded Physics
One of the primary goals of SPECFEM++ is to make adding new physics straightforward. The solver now supports elastic P-SV and SH waves, anisotropic media, poroelastic domains, and Cosserat media (which adds rotational degrees of freedom alongside displacement). Most significantly, 3D elastic isotropic simulation is now fully supported, with end-to-end integration tests confirming sample-by-sample agreement with SPECFEM3D Cartesian.
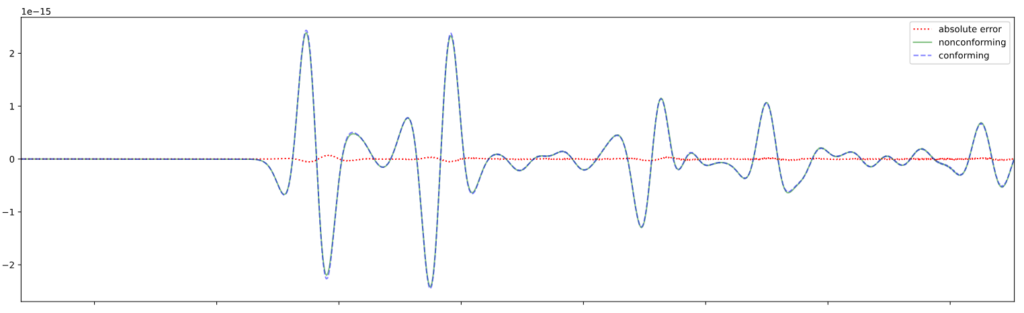
For problems with large contrasts in physical properties, non-conforming mesh support via a Discontinuous Galerkin approach allows different element sizes on either side of an interface. This enables the code to choose larger timesteps and drastically improve performance.
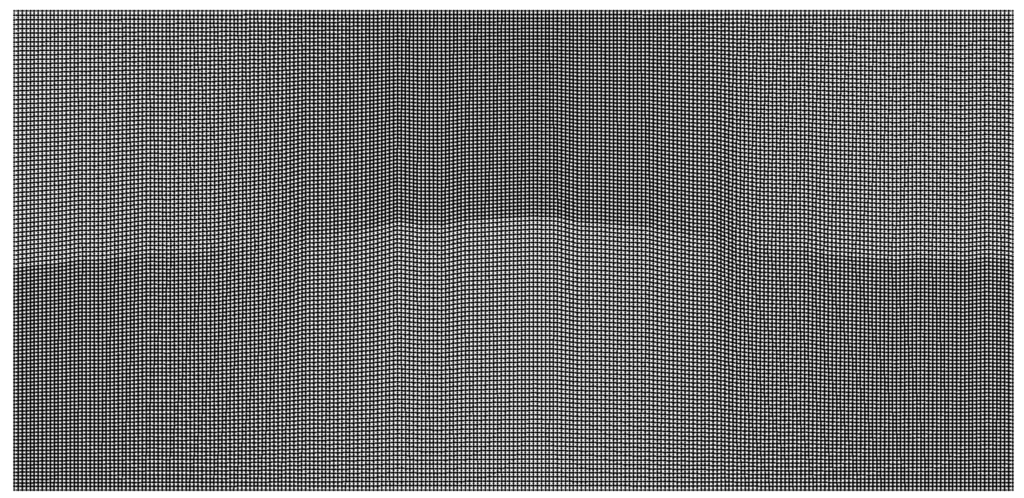
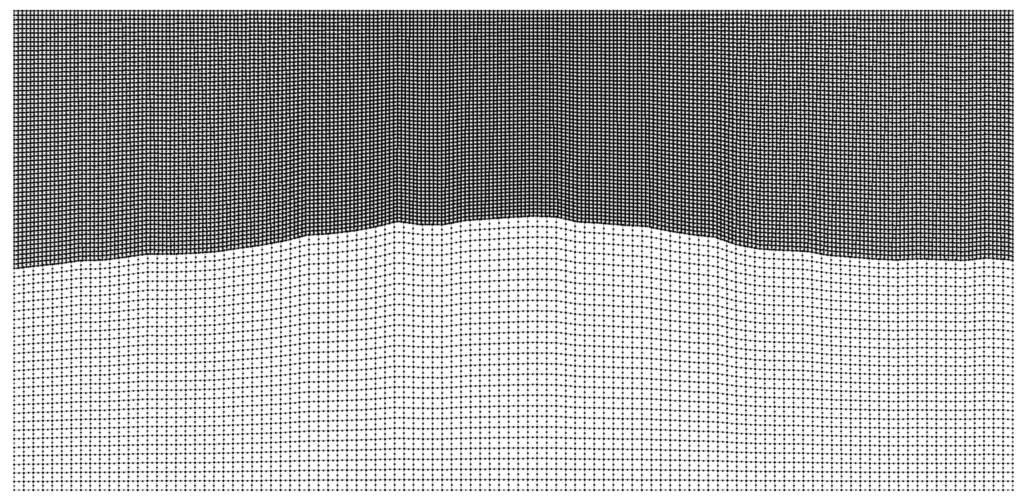


The conforming simulation is part of Pipatprathanporn et al., 2024 (DOI). Credit: Kentaro G. Hanson (G3, PACM).
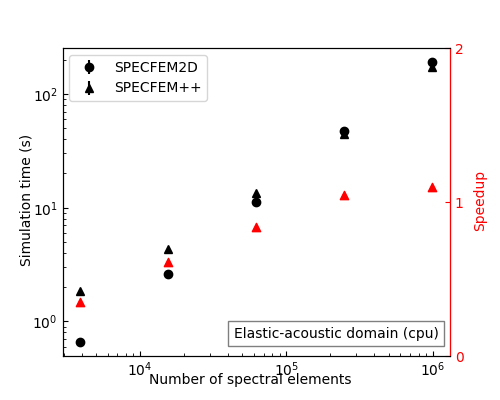
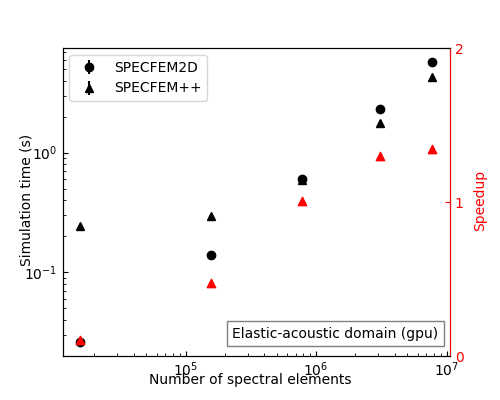
Performance on Par with — and Beyond — SPECFEM2D
After memory layout optimizations, SPECFEM++ now matches SPECFEM2D on CPU for elastic and acoustic problems. On GPU, it pulls ahead: for large elastic-acoustic domains (10M+ spectral elements), SPECFEM++ achieves up to 2× speedup over SPECFEM2D.
Real-World Benchmarks: Marmousi and Cosserat
Two showcases stand out. The Marmousi model cookbook runs wave propagation on a high-resolution CUBIT mesh derived from a classic seismic benchmark, and is a great stress test of the GPU backend. The Cosserat media simulation illustrates the rotational wavefield alongside displacement — something not possible in the original SPECFEM.

Tooling and Documentation
Nightly benchmarks run automatically via Jenkins on Intel Gold CPUs and NVIDIA H100 GPUs, with results on a live review dashboard. The API documentation now includes the actual implemented equations, and cookbooks cover everything from basic homogeneous media to non-conforming fluid-solid interfaces.
What’s Next
Active development is focused on MPI support, attenuation, and acoustic-elastic coupling — the remaining pieces needed for large-scale parallel 3D production runs.
SPECFEM++ is a community project with monthly developer meetings (first
Wednesday of each month, 12:00 PM Eastern sign up here). Documentation is at specfem2d-kokkos.readthedocs.io and the code is on GitHub.
Acknowledgements: The initial draft for this post was generated by Anthropic’s Claude Sonnet 4.6 using presentation slides and github release notes.